Detecting intraspecific diversity with eDNA
While environmental DNA (eDNA) approaches are well established for detecting species presence and biodiversity, little is known about the utility of these methods for gathering population-level genetic information. My research aims to develop and validate novel approaches to obtain intraspecific genetic information from eDNA samples. Using mesocosm experiments and field-based applications, I target mitochondrial and nuclear genetic markers to estimate population structure, genetic diversity, and species abundance of round gobies (Neogobius melanostomus) in eDNA samples. As part of a 2021 Graduate Research Fellowship with the Cooperative Institute for Great Lakes Research, I am also working with scientists at the USGS Great Lakes Science Center to develop novel eDNA-based metrics of species abundance based on genetic diversity.
Andres, K.J., Lodge, D.M., Andrés, J. (2023). Environmental DNA reveals the genetic diversity and population structure of an invasive species in the Laurentian Great Lakes. Proceedings of the National Academy of Sciences, 120 (37) e2307345120. DOI: 10.1073/pnas.2307345120
Andres, K.J., Lodge, D.M., Sethi, S.A., Andrés, J. (2023). Detecting and analysing intraspecific genetic variation with eDNA: From population genetics to species abundance. Molecular Ecology, 32(15), 4118–4132. DOI: 10.1111/mec.17031
Andres, K. J., Sethi, S. A., Lodge, D.M., & Andrés, J. (2021). Nuclear eDNA estimates population allele frequencies and abundance in experimental mesocosms and field samples. Molecular Ecology, 30, 685-697. doi: doi.org/10.1111/mec.15765
Andres, K.J., Lodge, D.M., Sethi, S.A., Andrés, J. (2023). Detecting and analysing intraspecific genetic variation with eDNA: From population genetics to species abundance. Molecular Ecology, 32(15), 4118–4132. DOI: 10.1111/mec.17031
Andres, K. J., Sethi, S. A., Lodge, D.M., & Andrés, J. (2021). Nuclear eDNA estimates population allele frequencies and abundance in experimental mesocosms and field samples. Molecular Ecology, 30, 685-697. doi: doi.org/10.1111/mec.15765
Multi-gear sampling to optimize biodiversity surveys
|
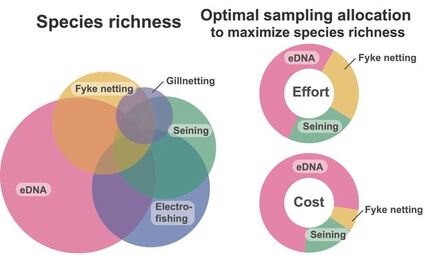
We evaluated the fish species richness observed in each sampling method (eDNA metabarcoding, electrofishing, fyke netting, gillnetting, and seining) individually and in combination to determine the allocation of effort and cost across sampling gear that provides the optimal approach for lake-wide species inventories. Optimal gear combination analyses revealed that detected species richness is maximized when most of the effort or budget is allocated to eDNA metabarcoding, with smaller allocations to seining and fyke netting.
|
Andres, K.J., Lambert, T., Lodge, D.M., Andrés, J., Jackson, R. (2023). Combining sampling gear to optimally inventory species highlights the efficiency of eDNA metabarcoding. Environmental DNA, 5, 146-157. DOI: 10.1002/edn3.366
Fish biodiversity in the Amazon River basin
The Amazon Basin contains the highest freshwater fish diversity on Earth, with over 2,500 described fish species residing within its rivers. However, due to the substantial size of rivers in the Amazon Basin, the sampling effort required to obtain biodiversity estimates using traditional sampling methods is impossible to achieve. With my research, I am testing the effectiveness of using environmental DNA (eDNA) approaches to characterize fish diversity and distributions in the western Amazon Basin, and examining the factors governing fish biodiversity in this ecosystem. This work will provide critical information on the structure of fish diversity in tropical rivers and will contribute to our understanding of how eDNA can be used to detect fish biodiversity in the Amazon Basin.
|
Morphological shifts to changing hydrology
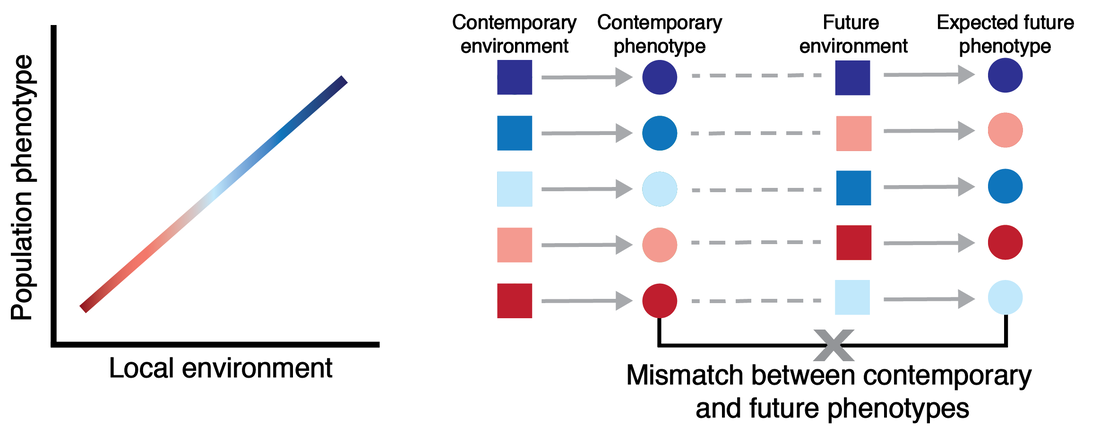
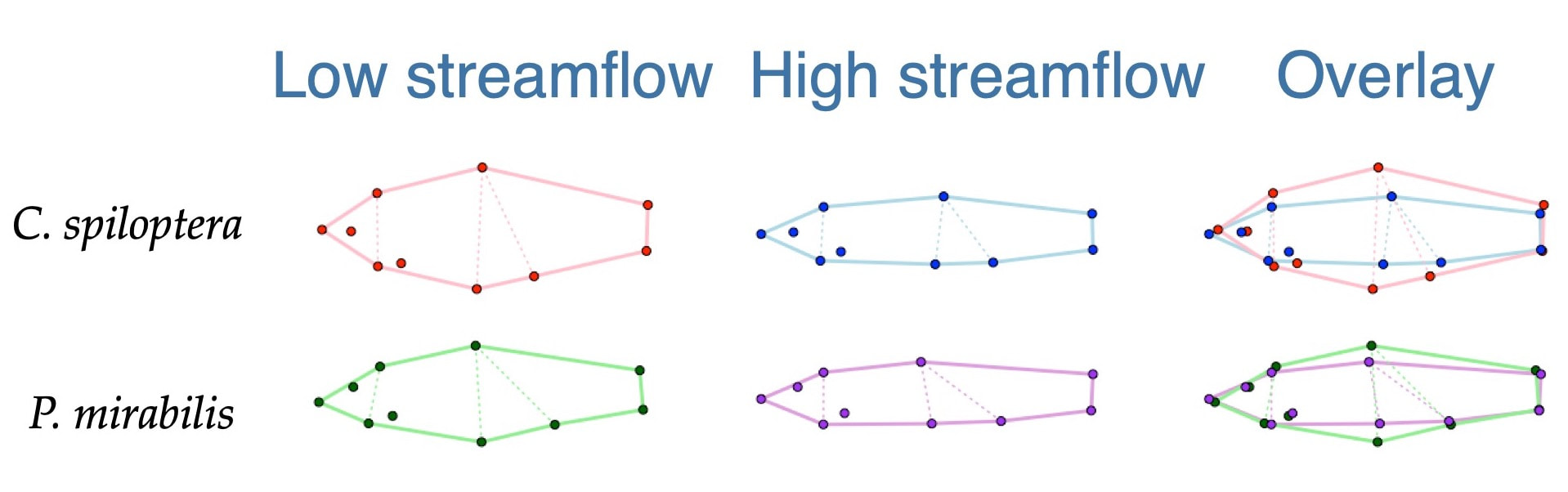
We examined patterns of population-level phenotypic changes expected under future climatic change using phenotype-environment associations between body shape and contemporary streamflow in minnow species. When phenotype-environment relationships were projected into future streamflow conditions, populations are not expected to respond uniformly across the species' ranges, and all but one species exhibited projected morphologies outside of the current range of morphological variation. These findings suggest local adaptation and spatial heterogeneity in environmental changes interact to influence variation in the degree of expected phenotypic responses to climate change at both the species and population level.
Andres, K.J., Chien, H., Knouft, J.H. Hydrology induces intraspecific variation in freshwater fish morphology under contemporary and future climate scenarios. (2019). Science of the Total Environment, 671, 421–430. DOI: 10.1016/j.scitotenv.2019.03.292
Andres, K.J., Chien, H., Knouft, J.H. Hydrology induces intraspecific variation in freshwater fish morphology under contemporary and future climate scenarios. (2019). Science of the Total Environment, 671, 421–430. DOI: 10.1016/j.scitotenv.2019.03.292